|
1/16/2024 0 Comments Salmonella on xld 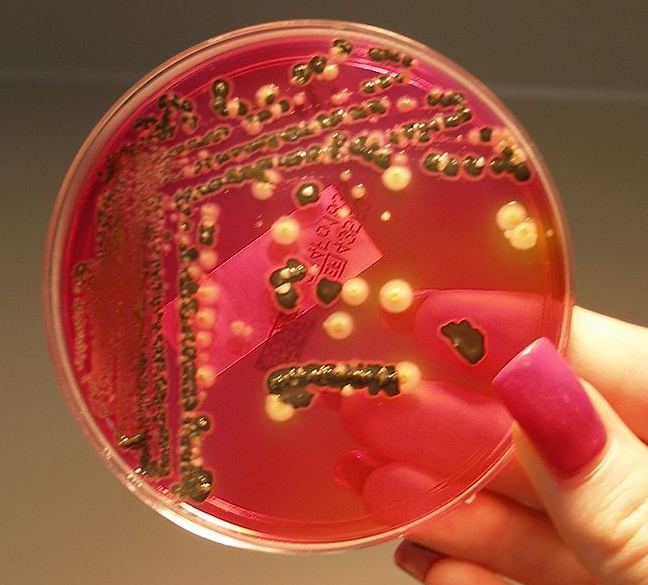
In many developed countries, AMR surveillance systems are replacing phenotypic antimicrobial susceptibility testing with WGS. Additionally, the selective culture and detection of bacteria with rare antimicrobial resistance (AMR) mechanisms is highly recommended. After phenotypic antibiotic susceptibility testing, genetic characterization can be done through the detection of targeted genes or through whole genome sequencing (WGS). Even though the broth microdilution method is preferred in many surveillance systems and in research, other less technology intensive methods of antibiotic susceptibility testing, such as disk diffusion, may also be used. After their isolation from samples, genus/species confirmation, and subtyping when necessary, the bacteria of interest are tested for susceptibility to a select number of antibiotics using a standard phenotyping method of choice. In most cases, surveillance systems of antibiotic resistant bacteria in food animals target pathogenic bacteria, such as Salmonella and Campylobacter as well as indicator bacteria, such as E. Monitoring of the emergence, spread, and changes in levels of antimicrobial resistant bacteria along the food chain is needed to inform and guide integrated strategies for combating antimicrobial resistance. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. In addition, sequencing data are accessible are accessible at the NCBI Sequence Reads Archive through the accession number PRJNA690823 The protocol can also be accessed at Protocols.io: dx.doi.org/10.17504/protocols.io.brjkm4kw.įunding: GHL,RM: This work was supported by the United States Agency for International Development, as part of the Feed the Future initiative, under the CGIAR Fund, award number BFS-G-11-00002. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.ĭata Availability: All relevant data are within the paper. 
Received: NovemAccepted: ApPublished: May 7, 2021Ĭopyright: © 2021 Manishimwe et al. PLoS ONE 16(5):Įditor: Iddya Karunasagar, Nitte University, INDIA The tested protocol is suited to detecting and estimating prevalence of antimicrobial resistance in bacteria isolated from food animal feces in resource-limited laboratories in the developing world.Ĭitation: Manishimwe R, Moncada PM, Bugarel M, Scott HM, Loneragan GH (2021) Antibiotic resistance among Escherichia coli and Salmonella isolated from dairy cattle feces in Texas. 
Whole genome sequencing showed that phenotypic profiles of antibiotic resistance detected were generally substantiated by genotypic profiles. coli isolates not susceptible to ceftriaxone, an AmpC phenotype was more common than an ESBL phenotype (29 versus 10 isolates, respectively). Sixteen Salmonella were isolated and only one demonstrated any resistance (i.e., single resistance to streptomycin). coli isolates recovered from MacConkey agar with ciprofloxacin, 77.3% and 54.5% were resistant to nalidixic acid and ciprofloxacin, respectively. coli recovered from MacConkey agar with cefotaxime, 56.8% were resistant ceftriaxone. coli isolated from MacConkey agar without antibiotics, the isolate-level prevalence of resistance to tetracycline was the highest (8.2%). Molecular methods were performed on selected bacterial isolates to identify and distinguish genetic determinants associated with the observed phenotypes. Bacterial isolates not susceptible to third-generation cephalosporins were tested for extended spectrum beta-lactamase (ESBL) phenotype using the combination disk test. All bacterial isolates were further tested for antibiotic susceptibility using disk diffusion. coli and Salmonella isolates not susceptible to third-generation cephalosporins and quinolones using selective media supplemented with cefotaxime (1.0 μg/mL) and ciprofloxacine (0.5 μg/mL), respectively. coli and Salmonella using selective media without antibiotic supplements. Using a predesigned protocol, fecal samples were collected to isolate non-type-specific E. The objective of this study was to develop and field-test a low cost protocol to estimate the isolate- and sample-level prevalence of resistance to critically important antibiotics among Escherichia coli and Salmonella isolated from dairy cattle feces. In several developing countries, studies on antimicrobial resistance among bacteria from food animals are rare mostly because of under-resourced laboratories. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
